About
I study how modern vision architectures learn, transfer, and fail when moved from natural images into biomedical microscopy, a domain where the standard assumptions of computer vision break down in informative ways.
My current work centres on a controlled benchmark comparing CNNs and Vision Transformers for single-cell classification on the LIVECell dataset, where I found that previously reported CNN superiority is an artefact of unmatched pretraining, and that standard optimisation recipes like layer-wise learning rate decay can unexpectedly hurt under domain shift.
I'm now building on these findings through domain-adaptive self-supervised learning (PhaseDINO), knowledge distillation, and cross-modality robustness testing.
More broadly, I'm interested in the questions that sit beneath the benchmarks: what makes one representation transfer better than another? Why do some training recipes fail outside their native domain? How do inductive biases interact with pretraining regime?
My path to these questions came through analytical philosophy, psychology, and neuroscience before arriving at machine learning, I've always been drawn to how systems form knowledge and why they sometimes get it wrong.
Selected results
From a controlled comparison of eleven architectures on LIVECell 8-class single-cell classification, with matched pretraining conditions across all models:
| Architecture | Macro F1 | Accuracy | Note |
|---|---|---|---|
| EfficientNet-B5 | 92.25% | 93.53% | Best single model |
| EVA-02 (ViT-B/14) | 90.51% | 92.26% | Best teacher for distillation |
| EVA-02 → EN-B0 | 91.01% | 92.67% | Cross-architecture distillation |
| EN-B5 → EN-B0 | 91.01% | 92.18% | Same-family distillation |
| 3×EVA-02 council → EN-B0 | 92.05% | 93.42% | ViT council distillation. Best distillation |
| 2xEVA-02 + EN-B5 council → EN-B0 | 91.41% | 92.86% | Mixed council distillation |
| ViT-B/16 | 89.93% | 91.94% | |
| ViT-S/8 | 90.61 | 92.20% | Matches EVA-02 at ¼ parameters, better than the larger ViT-B/16 due to smaller patches/finer tokenisation |
Layer-wise learning rate decay (LLRD), widely used for fine-tuning pretrained transformers, consistently hurts performance under ImageNet to LIVECell domain shift, a finding with implications for how standard NLP-derived training recipes are adopted in biomedical vision.
Gallery
Impressions from the bench, the microscope, and the training loop.
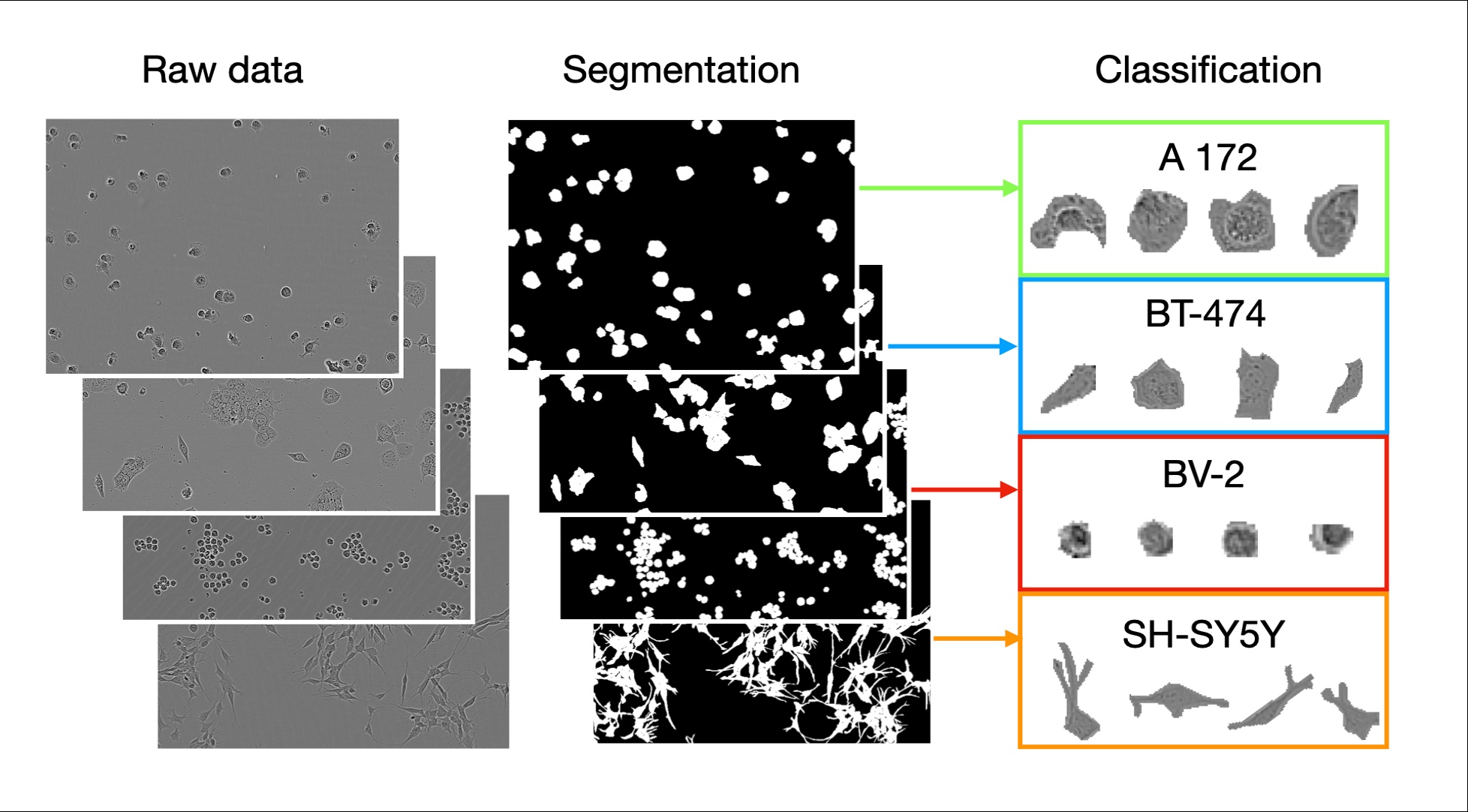 LIVECell pipeline: phase-contrast images → instance segmentation → single-cell classification
LIVECell pipeline: phase-contrast images → instance segmentation → single-cell classification
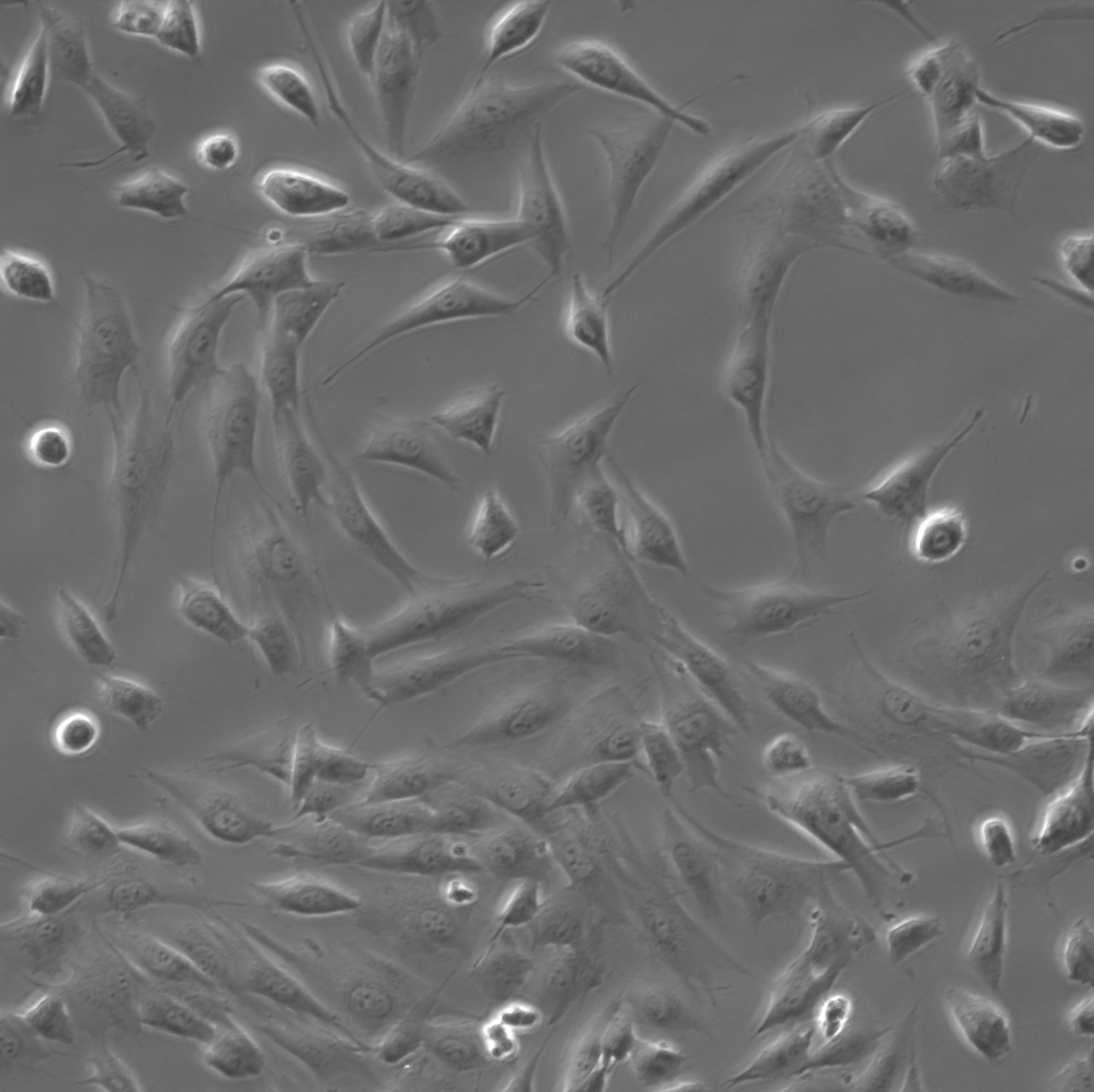 MG-63 osteosarcoma cells, phase-contrast 20×
MG-63 osteosarcoma cells, phase-contrast 20×
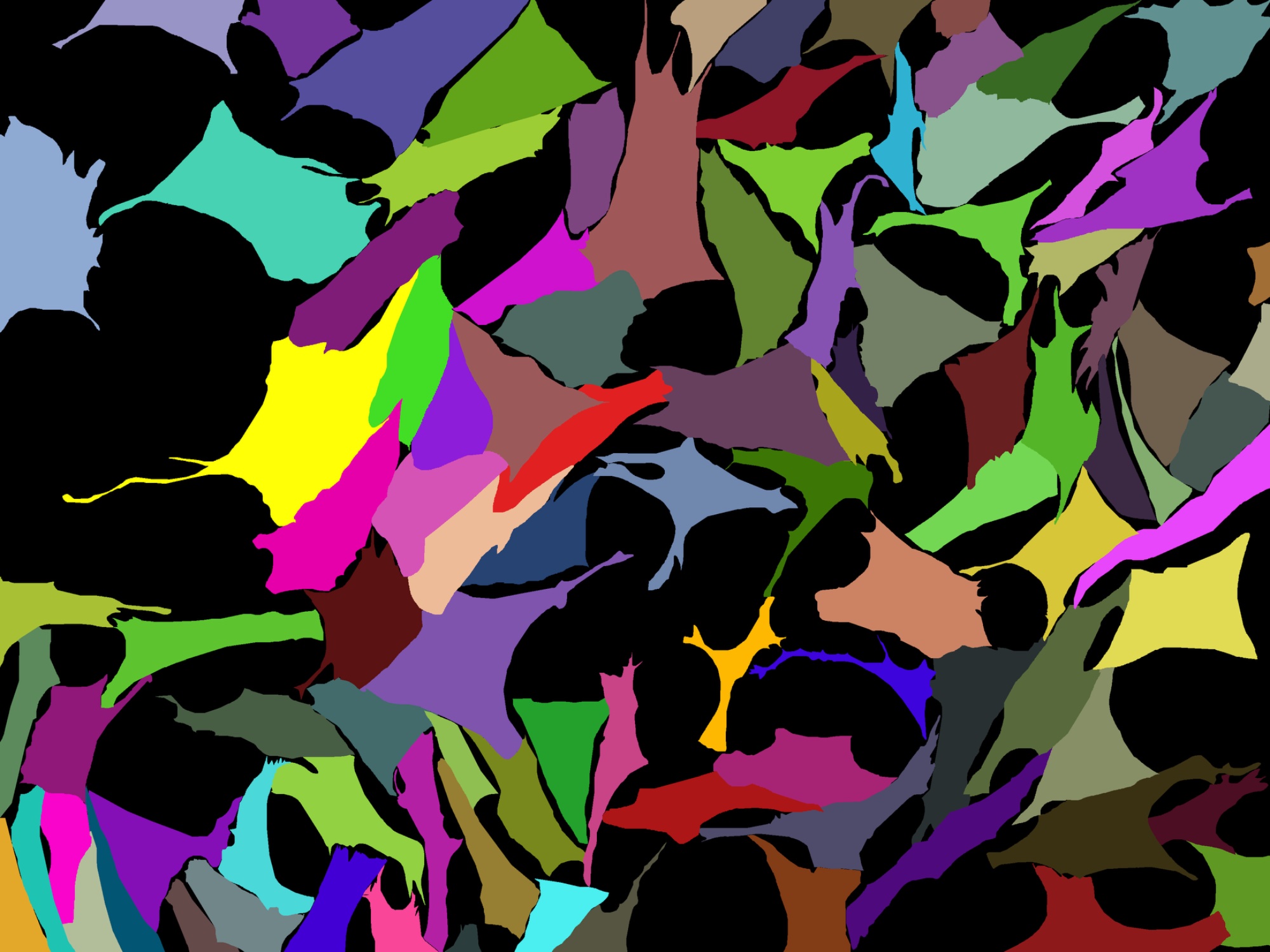 Instance segmentation mask, each colour is a single cell
Instance segmentation mask, each colour is a single cell
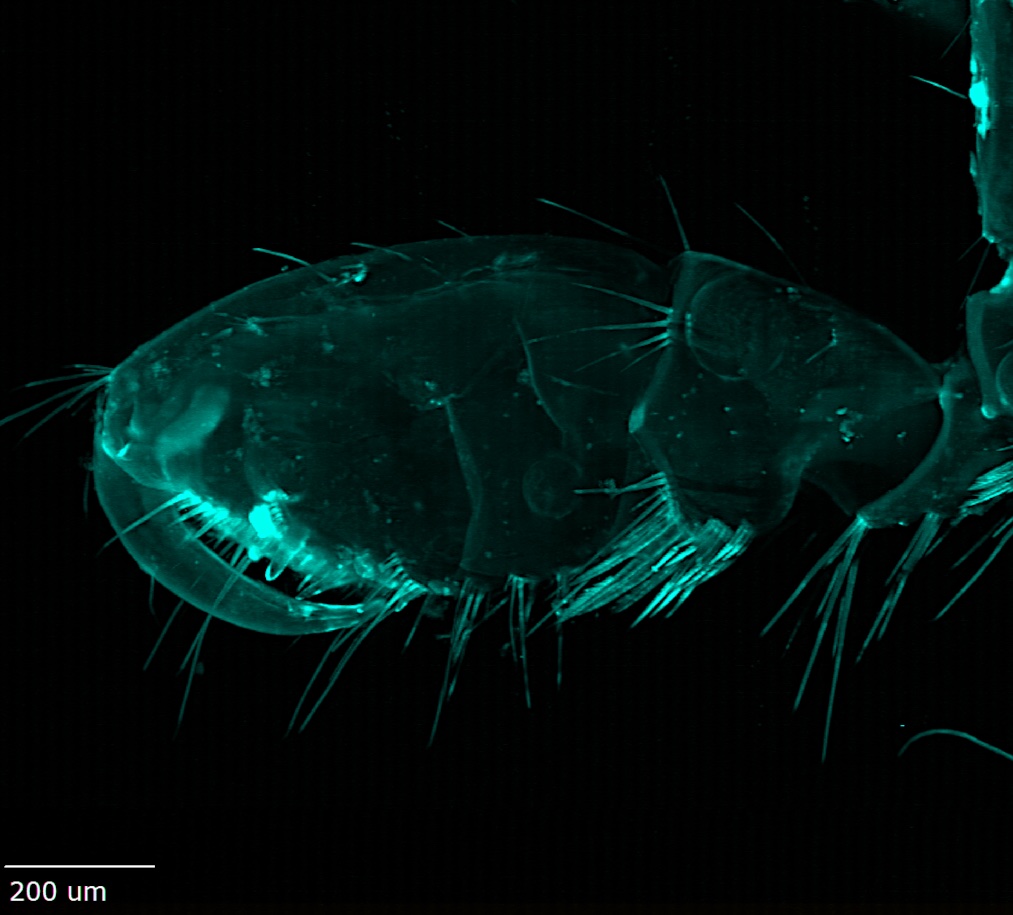 Amphipod copulatory claw (gnathopod), capturing autofluorescence, 200 µm scale bar
Amphipod copulatory claw (gnathopod), capturing autofluorescence, 200 µm scale bar
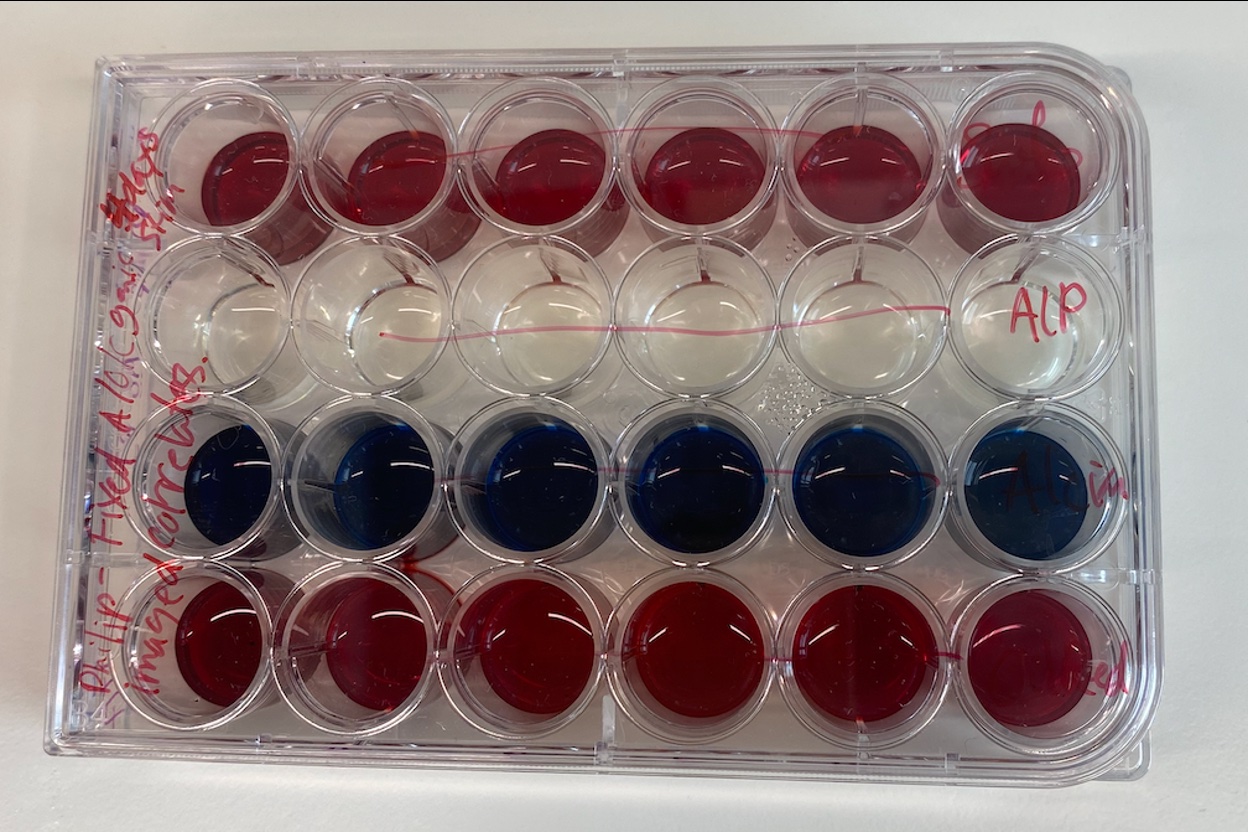 24-well plate: Alcian blue, oil red, ALP, Alizarin Red. Assessing differentiation of hMSCs.
24-well plate: Alcian blue, oil red, ALP, Alizarin Red. Assessing differentiation of hMSCs.
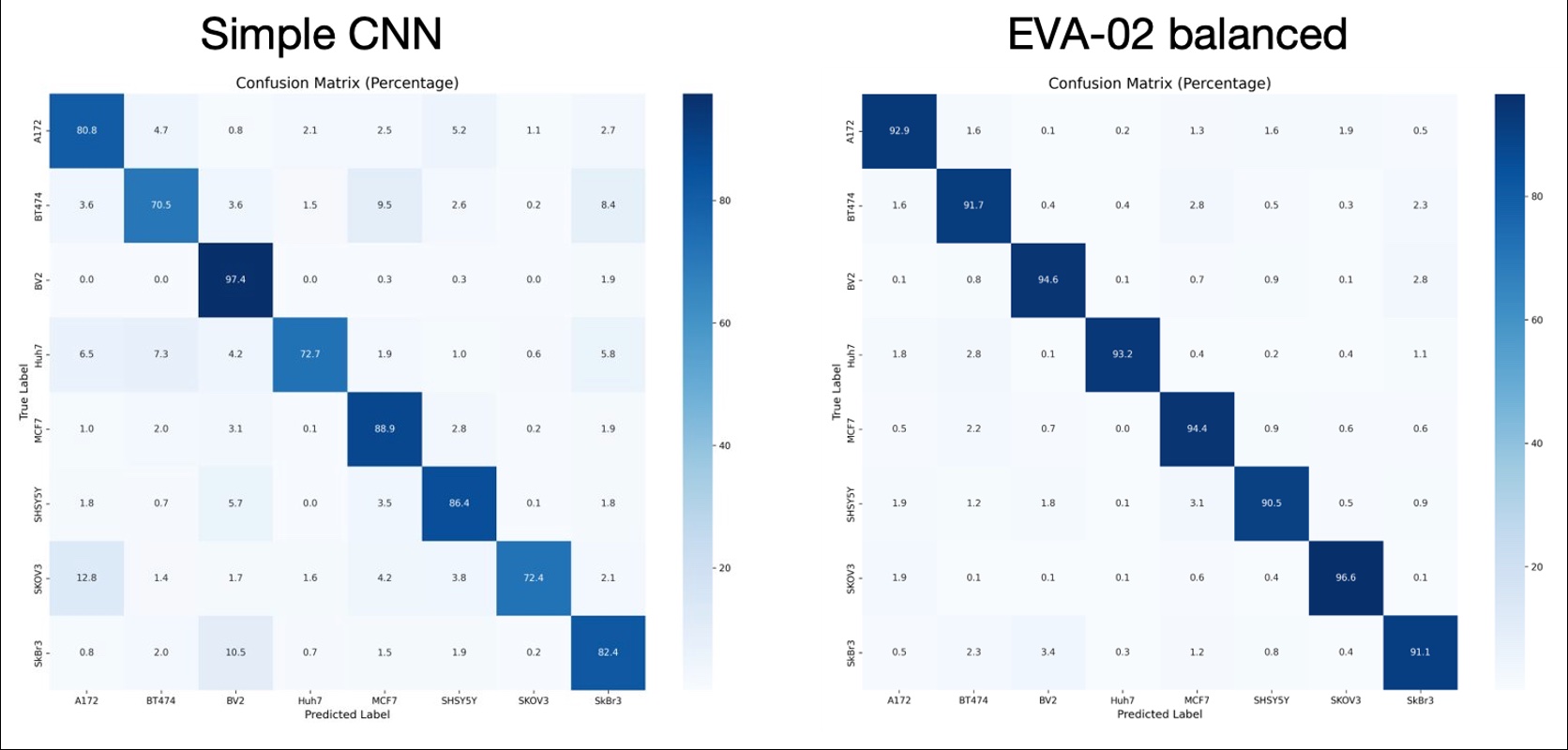 Custom CNN vs pretrained class-balanced EVA-02
Custom CNN vs pretrained class-balanced EVA-02
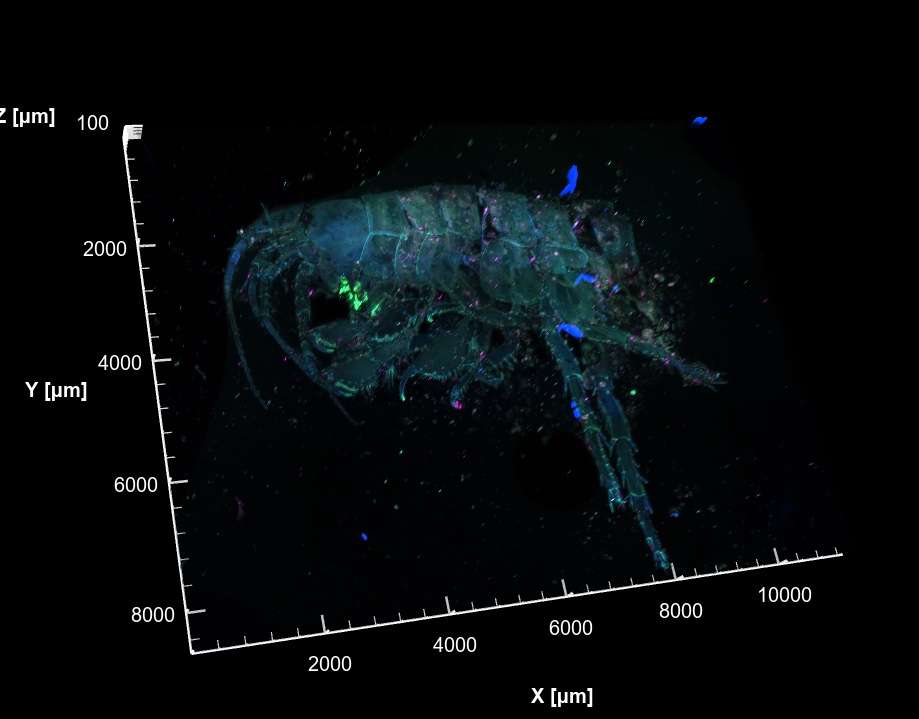 3D volume rendering of a whole amphipod, multi-channel fluorescence
3D volume rendering of a whole amphipod, multi-channel fluorescence
 574 segmented instances in a single field of view
574 segmented instances in a single field of view
Publications
Code
Controlled benchmark comparing CNN and Vision Transformer architectures for single-cell classification on LIVECell. Eleven architectures, pretrained and scratch conditions, multi-seed reproducibility.
Cross-architecture knowledge distillation. Temperature-scaled soft labels, combined hard/soft loss, council distillation. Didactic codebase with extensive inline documentation.
Validation tool for single-cell crops extracted from segmentation masks. Flags multi-cell crops, boundary truncation, size outliers, and mask coverage issues.